This vignette outlines a reproducible workflow for:
- loading packaged simulation data
- fitting an MFRM with flexible facets
- running diagnostics and residual PCA
- generating APA and visual summary outputs
- moving from fitted models into design simulation and fixed-calibration prediction
For a plot-first companion guide, see the separate
mfrmr-visual-diagnostics vignette.
For speed-sensitive work, a useful pattern is:
- start with
method = "JML"orquad_points = 7 - use
diagnose_mfrm(..., residual_pca = "none")for the first pass - reuse the same diagnostics object in downstream reports and plots
Load Data
library(mfrmr)
list_mfrmr_data()
#> [1] "example_core" "example_bias" "study1" "study2"
#> [5] "combined" "study1_itercal" "study2_itercal" "combined_itercal"
data("ej2021_study1", package = "mfrmr")
head(ej2021_study1)
#> Study Person Rater Criterion Score
#> 1 Study1 P001 R08 Global_Impression 4
#> 2 Study1 P001 R08 Linguistic_Realization 3
#> 3 Study1 P001 R08 Task_Fulfillment 3
#> 4 Study1 P001 R10 Global_Impression 4
#> 5 Study1 P001 R10 Linguistic_Realization 3
#> 6 Study1 P001 R10 Task_Fulfillment 2
study1_alt <- load_mfrmr_data("study1")
identical(names(ej2021_study1), names(study1_alt))
#> [1] TRUEMinimal Runnable Example
We start with the packaged example_core dataset. It is
intentionally compact, category-complete, and generated from a single
latent trait plus facet main effects so that help-page examples stay
fast without relying on undersized toy data. The same object is also
available via
data("mfrmr_example_core", package = "mfrmr"):
data("mfrmr_example_core", package = "mfrmr")
toy <- mfrmr_example_core
fit_toy <- fit_mfrm(
data = toy,
person = "Person",
facets = c("Rater", "Criterion"),
score = "Score",
method = "JML",
model = "RSM",
maxit = 15
)
#> Warning: Optimizer did not fully converge (code = 1). Consider increasing maxit
#> (current: 15) or relaxing reltol (current: 1e-06).
diag_toy <- diagnose_mfrm(fit_toy, residual_pca = "none")
summary(fit_toy)$overview
#> # A tibble: 1 × 11
#> Model Method N Persons Facets Categories LogLik AIC BIC Converged
#> <chr> <chr> <int> <int> <int> <dbl> <dbl> <dbl> <dbl> <lgl>
#> 1 RSM JMLE 768 48 2 4 -822. 1758. 2022. FALSE
#> # ℹ 1 more variable: Iterations <int>
summary(diag_toy)$overview
#> # A tibble: 1 × 8
#> Observations Persons Facets Categories Subsets ResidualPCA Method
#> <int> <int> <int> <int> <int> <chr> <chr>
#> 1 768 48 2 4 1 none JML
#> # ℹ 1 more variable: PrecisionTier <chr>
names(plot(fit_toy, draw = FALSE))
#> [1] "wright_map" "pathway_map"
#> [3] "category_characteristic_curves"Diagnostics and Reporting
t4_toy <- unexpected_response_table(
fit_toy,
diagnostics = diag_toy,
abs_z_min = 1.5,
prob_max = 0.4,
top_n = 10
)
t12_toy <- fair_average_table(fit_toy, diagnostics = diag_toy)
t13_toy <- bias_interaction_report(
estimate_bias(fit_toy, diag_toy,
facet_a = "Rater", facet_b = "Criterion",
max_iter = 2),
top_n = 10
)
class(summary(t4_toy))
#> [1] "summary.mfrm_bundle"
class(summary(t12_toy))
#> [1] "summary.mfrm_bundle"
class(summary(t13_toy))
#> [1] "summary.mfrm_bundle"
names(plot(t4_toy, draw = FALSE))
#> [1] "name" "data"
names(plot(t12_toy, draw = FALSE))
#> [1] "name" "data"
names(plot(t13_toy, draw = FALSE))
#> [1] "name" "data"Fit and Diagnose with Full Data
For a realistic analysis, we use the packaged Study 1 dataset:
fit <- fit_mfrm(
data = ej2021_study1,
person = "Person",
facets = c("Rater", "Criterion"),
score = "Score",
method = "MML",
model = "RSM",
quad_points = 7
)
diag <- diagnose_mfrm(
fit,
residual_pca = "none"
)
summary(fit)
#> Many-Facet Rasch Model Summary
#> Model: RSM | Method: MML
#> N: 1842 | Persons: 307 | Facets: 2 | Categories: 4
#> LogLik: -2102.737 | AIC: 4249.475 | BIC: 4370.884
#> Converged: Yes | Iterations: 80
#>
#> Facet overview
#> Facet Levels MeanEstimate SDEstimate MinEstimate MaxEstimate Span
#> Criterion 3 0 0.692 -0.799 0.430 1.229
#> Rater 18 0 0.665 -0.946 1.619 2.565
#>
#> Person measure distribution
#> Persons Mean SD Median Min Max Span MeanPosteriorSD
#> 307 0.414 0.812 0.439 -1.451 2.384 3.834 0.482
#>
#> Step parameter summary
#> Steps Min Max Span Monotonic
#> 3 -1.092 0.957 2.05 TRUE
#>
#> Most extreme facet levels (|estimate|)
#> Facet Level Estimate
#> Rater R13 1.619
#> Rater R08 -0.946
#> Rater R09 -0.918
#> Rater R06 0.886
#> Criterion Global_Impression -0.799
#>
#> Highest person measures
#> Person Estimate SD
#> P157 2.384 0.466
#> P239 2.346 0.543
#> P135 2.263 0.470
#> P018 2.121 0.612
#> P209 2.014 0.644
#>
#> Lowest person measures
#> Person Estimate SD
#> P136 -1.451 0.555
#> P173 -1.399 0.526
#> P159 -1.349 0.579
#> P048 -1.330 0.466
#> P089 -1.274 0.395
#>
#> Notes
#> - No immediate warnings from fit-level summary checks.
summary(diag)
#> Many-Facet Rasch Diagnostics Summary
#> Observations: 1842 | Persons: 307 | Facets: 2 | Categories: 4 | Subsets: 1
#> Residual PCA mode: none
#> Method: MML | Precision tier: model_based
#>
#> Overall fit
#> Infit Outfit InfitZSTD OutfitZSTD DF_Infit DF_Outfit
#> 0.811 0.786 -4.643 -7.029 1059.444 1842
#>
#> Precision basis
#> Method Converged PrecisionTier SupportsFormalInference HasFallbackSE
#> MML TRUE model_based TRUE FALSE
#> PersonSEBasis NonPersonSEBasis
#> Posterior SD (EAP) Observed information (MML)
#> CIBasis
#> Normal interval from model-based SE
#> ReliabilityBasis
#> Observed variance with model-based and fit-adjusted error bounds
#> HasFitAdjustedSE HasSamplePopulationCoverage
#> TRUE TRUE
#> RecommendedUse
#> Use for primary reporting of SE, CI, and reliability in this package.
#>
#> Facet precision and spread
#> Facet Levels Separation Strata Reliability RealSeparation RealStrata
#> Criterion 3 14.910 20.214 0.996 14.910 20.214
#> Person 307 1.322 2.096 0.636 1.225 1.967
#> Rater 18 3.118 4.490 0.907 3.107 4.476
#> RealReliability MeanInfit MeanOutfit
#> 0.996 0.810 0.786
#> 0.600 0.798 0.786
#> 0.906 0.813 0.786
#>
#> Largest |ZSTD| rows
#> Facet Level Infit Outfit InfitZSTD OutfitZSTD AbsZ
#> Criterion Global_Impression 0.798 0.744 -2.596 -4.922 4.922
#> Rater R08 0.702 0.660 -2.439 -4.107 4.107
#> Criterion Linguistic_Realization 0.802 0.797 -2.918 -3.817 3.817
#> Person P020 0.027 0.026 -2.768 -3.464 3.464
#> Criterion Task_Fulfillment 0.829 0.816 -2.488 -3.422 3.422
#> Rater R10 0.737 0.726 -2.189 -2.939 2.939
#> Person P203 0.041 0.073 -2.497 -2.832 2.832
#> Person P098 2.314 3.361 1.539 2.780 2.780
#> Rater R05 0.744 0.749 -1.923 -2.535 2.535
#> Person P056 0.075 0.125 -1.727 -2.405 2.405
#>
#> Flag counts
#> Metric Count
#> Unexpected responses 100
#> Flagged displacement levels 40
#> Interaction rows 20
#> Inter-rater pairs 153
#>
#> Notes
#> - Unexpected responses were flagged under current thresholds.
#> - SE/ModelSE, CI, and reliability conventions depend on the estimation path; see diagnostics$approximation_notes for MML-vs-JML details.If you need dimensionality evidence for a final report, you can add residual PCA after the initial diagnostic pass:
diag_pca <- diagnose_mfrm(
fit,
residual_pca = "both",
pca_max_factors = 6
)
summary(diag_pca)
#> Many-Facet Rasch Diagnostics Summary
#> Observations: 1842 | Persons: 307 | Facets: 2 | Categories: 4 | Subsets: 1
#> Residual PCA mode: both
#> Method: MML | Precision tier: model_based
#>
#> Overall fit
#> Infit Outfit InfitZSTD OutfitZSTD DF_Infit DF_Outfit
#> 0.811 0.786 -4.643 -7.029 1059.444 1842
#>
#> Precision basis
#> Method Converged PrecisionTier SupportsFormalInference HasFallbackSE
#> MML TRUE model_based TRUE FALSE
#> PersonSEBasis NonPersonSEBasis
#> Posterior SD (EAP) Observed information (MML)
#> CIBasis
#> Normal interval from model-based SE
#> ReliabilityBasis
#> Observed variance with model-based and fit-adjusted error bounds
#> HasFitAdjustedSE HasSamplePopulationCoverage
#> TRUE TRUE
#> RecommendedUse
#> Use for primary reporting of SE, CI, and reliability in this package.
#>
#> Facet precision and spread
#> Facet Levels Separation Strata Reliability RealSeparation RealStrata
#> Criterion 3 14.910 20.214 0.996 14.910 20.214
#> Person 307 1.322 2.096 0.636 1.225 1.967
#> Rater 18 3.118 4.490 0.907 3.107 4.476
#> RealReliability MeanInfit MeanOutfit
#> 0.996 0.810 0.786
#> 0.600 0.798 0.786
#> 0.906 0.813 0.786
#>
#> Largest |ZSTD| rows
#> Facet Level Infit Outfit InfitZSTD OutfitZSTD AbsZ
#> Criterion Global_Impression 0.798 0.744 -2.596 -4.922 4.922
#> Rater R08 0.702 0.660 -2.439 -4.107 4.107
#> Criterion Linguistic_Realization 0.802 0.797 -2.918 -3.817 3.817
#> Person P020 0.027 0.026 -2.768 -3.464 3.464
#> Criterion Task_Fulfillment 0.829 0.816 -2.488 -3.422 3.422
#> Rater R10 0.737 0.726 -2.189 -2.939 2.939
#> Person P203 0.041 0.073 -2.497 -2.832 2.832
#> Person P098 2.314 3.361 1.539 2.780 2.780
#> Rater R05 0.744 0.749 -1.923 -2.535 2.535
#> Person P056 0.075 0.125 -1.727 -2.405 2.405
#>
#> Flag counts
#> Metric Count
#> Unexpected responses 100
#> Flagged displacement levels 40
#> Interaction rows 20
#> Inter-rater pairs 153
#>
#> Notes
#> - Unexpected responses were flagged under current thresholds.
#> - SE/ModelSE, CI, and reliability conventions depend on the estimation path; see diagnostics$approximation_notes for MML-vs-JML details.Residual PCA and Reporting
pca <- analyze_residual_pca(diag_pca, mode = "both")
plot_residual_pca(pca, mode = "overall", plot_type = "scree")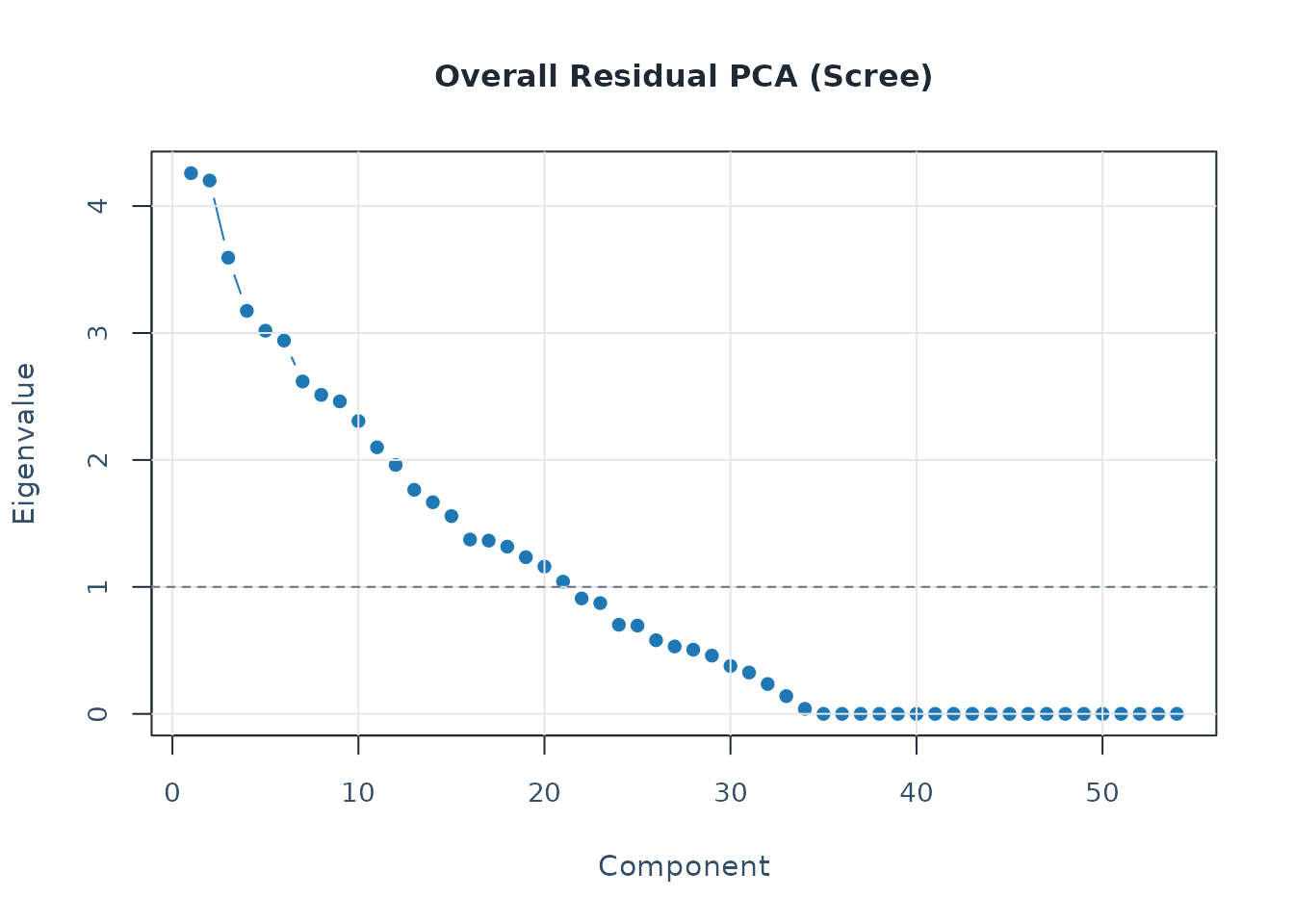
data("mfrmr_example_bias", package = "mfrmr")
bias_df <- mfrmr_example_bias
fit_bias <- fit_mfrm(
bias_df,
person = "Person",
facets = c("Rater", "Criterion"),
score = "Score",
method = "MML",
model = "RSM",
quad_points = 7
)
diag_bias <- diagnose_mfrm(fit_bias, residual_pca = "none")
bias <- estimate_bias(fit_bias, diag_bias, facet_a = "Rater", facet_b = "Criterion")
fixed <- build_fixed_reports(bias)
apa <- build_apa_outputs(fit_bias, diag_bias, bias_results = bias)
mfrm_threshold_profiles()
#> $profiles
#> $profiles$strict
#> $profiles$strict$n_obs_min
#> [1] 200
#>
#> $profiles$strict$n_person_min
#> [1] 50
#>
#> $profiles$strict$low_cat_min
#> [1] 15
#>
#> $profiles$strict$min_facet_levels
#> [1] 4
#>
#> $profiles$strict$misfit_ratio_warn
#> [1] 0.08
#>
#> $profiles$strict$missing_fit_ratio_warn
#> [1] 0.15
#>
#> $profiles$strict$zstd2_ratio_warn
#> [1] 0.08
#>
#> $profiles$strict$zstd3_ratio_warn
#> [1] 0.03
#>
#> $profiles$strict$expected_var_min
#> [1] 0.3
#>
#> $profiles$strict$pca_first_eigen_warn
#> [1] 1.5
#>
#> $profiles$strict$pca_first_prop_warn
#> [1] 0.1
#>
#>
#> $profiles$standard
#> $profiles$standard$n_obs_min
#> [1] 100
#>
#> $profiles$standard$n_person_min
#> [1] 30
#>
#> $profiles$standard$low_cat_min
#> [1] 10
#>
#> $profiles$standard$min_facet_levels
#> [1] 3
#>
#> $profiles$standard$misfit_ratio_warn
#> [1] 0.1
#>
#> $profiles$standard$missing_fit_ratio_warn
#> [1] 0.2
#>
#> $profiles$standard$zstd2_ratio_warn
#> [1] 0.1
#>
#> $profiles$standard$zstd3_ratio_warn
#> [1] 0.05
#>
#> $profiles$standard$expected_var_min
#> [1] 0.2
#>
#> $profiles$standard$pca_first_eigen_warn
#> [1] 2
#>
#> $profiles$standard$pca_first_prop_warn
#> [1] 0.1
#>
#>
#> $profiles$lenient
#> $profiles$lenient$n_obs_min
#> [1] 60
#>
#> $profiles$lenient$n_person_min
#> [1] 20
#>
#> $profiles$lenient$low_cat_min
#> [1] 5
#>
#> $profiles$lenient$min_facet_levels
#> [1] 2
#>
#> $profiles$lenient$misfit_ratio_warn
#> [1] 0.15
#>
#> $profiles$lenient$missing_fit_ratio_warn
#> [1] 0.3
#>
#> $profiles$lenient$zstd2_ratio_warn
#> [1] 0.15
#>
#> $profiles$lenient$zstd3_ratio_warn
#> [1] 0.08
#>
#> $profiles$lenient$expected_var_min
#> [1] 0.1
#>
#> $profiles$lenient$pca_first_eigen_warn
#> [1] 3
#>
#> $profiles$lenient$pca_first_prop_warn
#> [1] 0.2
#>
#>
#>
#> $pca_reference_bands
#> $pca_reference_bands$eigenvalue
#> critical_minimum caution common strong
#> 1.4 1.5 2.0 3.0
#>
#> $pca_reference_bands$proportion
#> minor caution strong
#> 0.05 0.10 0.20
#>
#>
#> attr(,"class")
#> [1] "mfrm_threshold_profiles" "list"
vis <- build_visual_summaries(fit_bias, diag_bias, threshold_profile = "standard")
vis$warning_map$residual_pca_overall
#> [1] "Threshold profile: standard (PC1 EV >= 2.0, variance >= 10%)."
#> [2] "Heuristic reference bands: EV >= 1.4 (critical minimum), >= 1.5 (caution), >= 2.0 (common), >= 3.0 (strong); variance >= 5% (minor), >= 10% (caution), >= 20% (strong)."
#> [3] "Current exploratory PC1 checks: EV>=1.5:Y, EV>=2.0:Y, EV>=3.0:Y, Var>=10%:Y, Var>=20%:Y."
#> [4] "Overall residual PCA PC1 exceeds the current heuristic eigenvalue band (3.22)."
#> [5] "Overall residual PCA PC1 explains 20.1% variance."The same example_bias dataset also carries a
Group variable so DIF-oriented examples can show a non-null
pattern instead of a fully clean result. It can be loaded either with
load_mfrmr_data("example_bias") or
data("mfrmr_example_bias", package = "mfrmr").
Human-Readable Reporting API
spec <- specifications_report(fit, title = "Study run")
data_qc <- data_quality_report(
fit,
data = ej2021_study1,
person = "Person",
facets = c("Rater", "Criterion"),
score = "Score"
)
iter <- estimation_iteration_report(fit, max_iter = 8)
subset_rep <- subset_connectivity_report(fit, diagnostics = diag)
facet_stats <- facet_statistics_report(fit, diagnostics = diag)
cat_structure <- category_structure_report(fit, diagnostics = diag)
cat_curves <- category_curves_report(fit, theta_points = 101)
bias_rep <- bias_interaction_report(bias, top_n = 20)
plot_bias_interaction(bias_rep, plot = "scatter")
Design Simulation and Prediction
The package also supports a separate simulation/prediction layer. The key distinction is:
-
evaluate_mfrm_design()andpredict_mfrm_population()are design-level helpers that summarize expected operating characteristics under an explicit simulation specification. -
predict_mfrm_units()andsample_mfrm_plausible_values()score future or partially observed persons under a fixedMMLcalibration.
sim_spec <- build_mfrm_sim_spec(
n_person = 30,
n_rater = 4,
n_criterion = 4,
raters_per_person = 2,
assignment = "rotating"
)
pred_pop <- predict_mfrm_population(
sim_spec = sim_spec,
reps = 2,
maxit = 10,
seed = 1
)
#> Warning: Optimizer did not fully converge (code = 1). Consider increasing maxit
#> (current: 10) or relaxing reltol (current: 1e-06).
#> Warning: Optimizer did not fully converge (code = 1). Consider increasing maxit
#> (current: 10) or relaxing reltol (current: 1e-06).
summary(pred_pop)$forecast[, c("Facet", "MeanSeparation", "McseSeparation")]
#> # A tibble: 3 × 3
#> Facet MeanSeparation McseSeparation
#> <chr> <dbl> <dbl>
#> 1 Criterion 1.87 0.076
#> 2 Person 2.03 0.027
#> 3 Rater 0.728 0.728
keep_people <- unique(toy$Person)[1:18]
toy_mml <- suppressWarnings(
fit_mfrm(
toy[toy$Person %in% keep_people, , drop = FALSE],
person = "Person",
facets = c("Rater", "Criterion"),
score = "Score",
method = "MML",
quad_points = 5,
maxit = 15
)
)
new_units <- data.frame(
Person = c("NEW01", "NEW01"),
Rater = unique(toy$Rater)[1],
Criterion = unique(toy$Criterion)[1:2],
Score = c(2, 3)
)
pred_units <- predict_mfrm_units(toy_mml, new_units, n_draws = 0)
pv_units <- sample_mfrm_plausible_values(toy_mml, new_units, n_draws = 2, seed = 1)
summary(pred_units)$estimates[, c("Person", "Estimate", "Lower", "Upper")]
#> # A tibble: 1 × 4
#> Person Estimate Lower Upper
#> <chr> <dbl> <dbl> <dbl>
#> 1 NEW01 -0.097 -1.36 1.36
summary(pv_units)$draw_summary[, c("Person", "Draws", "MeanValue")]
#> # A tibble: 1 × 3
#> Person Draws MeanValue
#> <chr> <dbl> <dbl>
#> 1 NEW01 2 0